The SBML Test Suite is a conformance testing system for SBML. It can be used by developers to test their software’s implementation of SBML support; it can also be used by modelers to assess the correctness and completeness of SBML support in a given software system. The collection of test cases includes both syntactic and semantic tests.
More about the SBML Test Suite
Computational modeling has become a crucial aspect of biological research, and SBML has become the de facto standard open format for exchanging models between software tools in systems biology. However, implementing full support for SBML (especially for numerical simulations of models described by SBML) is a complex undertaking. The goal of the SBML Test Suite is to help developers implement correct SBML support, and help users evaluate the completeness of support in a given software tool.
The SBML Test Suite consists of several parts:
- A collection of SBML models, together with expected results, that can be used to test interpretation of SBML files. The tests are grouped into three subsets: semantic (for deterministic simulation behavior), stochastic (for stochastic simulation behavior), and syntactic (for basic parsing). Each suite of test cases can be downloaded separately as complete archives.
- The SBML Test Runner, a standalone application that can be used to run an application against the collection of semantic test cases, and report successes and failures. The image below shows an example run from the SBML Test Runner:
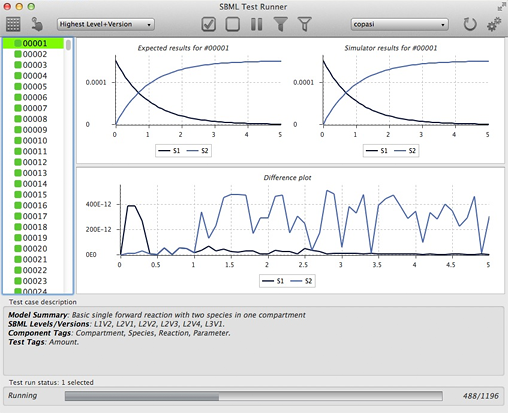
- An online database of test results provided by software authors for their SBML-compatible software packages.
SBML Test Suite release DOIs
We provide Zenodo DOIs for specific releases, so that you can cite the exact version of the software that you use:
- SBML Test Suite 3.3.0 ⇒ 10.5281/zenodo.1112521
Getting started
Please visit the SBML Test Suite documentation pages for information about how to get started with using the test suite.
Acknowledgments
Funding for the development of the core of the SBML Test Suite came from National Institute of General Medical Sciences grants R01 GM070923 and R01 GM077671 (Principal Investigator: Michael Hucka) and BBSRC grants BEP 17042, BBS/B/16550, and BBC0082001 (Principal Investigator: Darren J. Wilkinson). Additional support for this and other projects of the SBML Team have been provided by the following organizations since the year 2000: the International Joint Research Program of NEDO (Japan); the JST ERATO-SORST Program (Japan); the Japanese Ministry of Agriculture; the Japanese Ministry of Education, Culture, Sports, Science and Technology; the BBSRC e-Science Initiative (UK); the DARPA IPTO Bio-Computation Program (USA); the Army Research Office’s Institute for Collaborative Biotechnologies (USA); the Air Force Office of Scientific Research (USA); the California Institute of Technology (USA); the University of Hertfordshire (UK); the Molecular Sciences Institute (USA); the Systems Biology Institute (Japan); and the University of Keio (Japan).
The SBML Test Suite has been in development for many years, and many people have contributed in various ways large and small. The following are the primary authors of different parts:
-
Semantic test cases:
Sarah M. Keatinga,b, Lucian P. Smithb,c, Bruce Shapiro b, Michael Huckab, Frank T. Bergmannd, Brett Olivierg, Andrew Finneyb
-
Stochastic test cases:
Thomas W. Evanse, Colin S. Gillespief, Darren J. Wilkinsonf, Lucian P. Smithb,c
-
Syntactic test cases:
Sarah M. Keatinga,b, Lucian P. Smithb,c
-
SBML Test Runners:
Frank T. Bergmannd, Michael Huckab, Kimberly Begley
-
Online SBML Test Suite Database:
Frank T. Bergmannd
Additional contributions are gratefully acknowledged from Ralph Gaugesh, Chris Myersi, Akira Funahashij, Andreas Drägerk, Roland Kellerk, Fedor Kolpakovl, Stanley Guc, Matthias Königm, and others in the SBML community.
Institutions:
a EMBL-EBI, Hinxton, Cambridgeshire, UK
b California Institute of Technology, Pasadena, CA, US
c University of Washington, Seattle, WA, US
d University of Heidelberg, Heidelberg, DE
e University of Liverpool, Liverpool, UK
f Newcastle University, Newcastle, UK
g Vrije Universiteit Amsterdam
h Heidelberg Institute for Theoretical Studies, Heidelberg, DE
i University of Utah, Salt Lake City, UT, US
j Systems Biology Institute, Tokyo, JP
k University of Tuebingen, Tübingen, DE
l Institute of Systems Biology, Novosibirsk, RU
m Institute for Theoretical Biology, Humboldt Universität zu Berlin, DE
